-Search query
-Search result
Showing 1 - 50 of 8,695 items for (author: liu & h)
EMDB-60628: 
Carazolol-activated human beta3 adrenergic receptor
Method: single particle / : Zheng S, Zhang S, Dai S, Chen K, Gao K, Lin B, Liu X
EMDB-60629: 
Epinephrine-activated human beta3 adrenergic receptor
Method: single particle / : Zheng S, Zhang S, Dai S, Chen K, Gao K, Lin B, Liu X

EMDB-31367: 
Structure of mumps virus nucleoprotein without C-arm
Method: helical / : Shen Q, Shan H, Zhang N, Qin Y
EMDB-37217: 
Cryo-EM structure of human NADK tetramer
Method: single particle / : Zhang P, Hu M, Liu Z
EMDB-36364: 
G protein-coupled receptor 1
Method: single particle / : Liu A, Liu Y, Chen G, Ye F
EMDB-37197: 
The cryo-EM structure of type3 amyloid beta 42 fibril.
Method: helical / : Zhao QY, Tao YQ, Liu C, Li D

EMDB-37200: 
The cryo-EM structure of AV-45 bound type3 amyloid beta 42 fibril.
Method: helical / : Zhao QY, Tao YQ, Liu C, Li D

EMDB-37199: 
The cryo-EM structure of type1 amyloid beta 42 fibril in AD3.
Method: helical / : Zhao QY, Tao YQ, Liu C, Li D

EMDB-37195: 
The cryo-EM structure of AV-45 bound type1 amyloid beta 42 fibril.
Method: helical / : Zhao QY, Tao YQ, Liu C, Li D

EMDB-60099: 
SARS-CoV-2 spike trimer (6P) in complex with two R1-26 Fabs
Method: single particle / : Yan Q, Gao X, Liu B, Hou R, He P, Li Z, Chen Q, Wang J, He J, Chen L, Zhao J, Xiong X

EMDB-60100: 
SARS-CoV-2 spike trimer (6P) in complex with three R1-26 Fabs
Method: single particle / : Yan Q, Gao X, Liu B, Hou R, He P, Li Z, Chen Q, Wang J, He J, Chen L, Zhao J, Xiong X

EMDB-60101: 
SARS-CoV-2 spike trimer (6P) in complex with R1-26 Fab, head-to-head aggregate
Method: single particle / : Yan Q, Gao X, Liu B, Hou R, He P, Li Z, Chen Q, Wang J, He J, Chen L, Zhao J, Xiong X

EMDB-60102: 
SARS-CoV-2 spike trimer (6P) in complex with R1-26 Fab, focused refinement of RBD-Fab region
Method: single particle / : Yan Q, Gao X, Liu B, Hou R, He P, Li Z, Chen Q, Wang J, He J, Chen L, Zhao J, Xiong X

EMDB-60103: 
SARS-CoV-2 spike trimer (6P) in complex with two H18 Fabs
Method: single particle / : Yan Q, Gao X, Liu B, Hou R, He P, Li Z, Chen Q, Wang J, He J, Chen L, Zhao J, Xiong X

EMDB-60104: 
SARS-CoV-2 spike trimer (6P) in complex with three H18 Fabs
Method: single particle / : Yan Q, Gao X, Liu B, Hou R, He P, Li Z, Chen Q, Wang J, He J, Chen L, Zhao J, Xiong X

EMDB-60105: 
SARS-CoV-2 spike trimer (6P) in complex with three H18 Fabs, head-to-head aggregate (C1 symmetry)
Method: single particle / : Yan Q, Gao X, Liu B, Hou R, He P, Li Z, Chen Q, Wang J, He J, Chen L, Zhao J, Xiong X

EMDB-60106: 
SARS-CoV-2 spike trimer (6P) in complex with three H18 Fabs, head-to-head aggregate (C3 symmetry)
Method: single particle / : Yan Q, Gao X, Liu B, Hou R, He P, Li Z, Chen Q, Wang J, He J, Chen L, Zhao J, Xiong X

EMDB-60107: 
SARS-CoV-2 spike trimer (6P) in complex with two H18 and two R1-32 Fabs
Method: single particle / : Yan Q, Gao X, Liu B, Hou R, He P, Li Z, Chen Q, Wang J, He J, Chen L, Zhao J, Xiong X

EMDB-60108: 
SARS-CoV-2 spike trimer (6P) in complex with three H18 and three R1-32 Fabs
Method: single particle / : Yan Q, Gao X, Liu B, Hou R, He P, Li Z, Chen Q, Wang J, He J, Chen L, Zhao J, Xiong X

EMDB-60109: 
SARS-CoV-2 spike trimer (6P) in complex with three H18 and three R1-32 Fabs (one RBD rotated)
Method: single particle / : Yan Q, Gao X, Liu B, Hou R, He P, Li Z, Chen Q, Wang J, He J, Chen L, Zhao J, Xiong X

EMDB-60110: 
SARS-CoV-2 S1 in complex with H18 and R1-32 Fab
Method: single particle / : Yan Q, Gao X, Liu B, Hou R, He P, Li Z, Chen Q, Wang J, He J, Chen L, Zhao J, Xiong X

EMDB-60111: 
Dimer of SARS-CoV-2 S1 in complex with H18 and R1-32 Fabs
Method: single particle / : Yan Q, Gao X, Liu B, Hou R, He P, Li Z, Chen Q, Wang J, He J, Chen L, Zhao J, Xiong X

EMDB-37004: 
Oligonucleosome 0 DMSO
Method: subtomogram averaging / : Chen JK, Cai S, Liu T, Ruan W, Ng CT, Shi J, Surana U, Gan L

EMDB-37005: 
Oligonucleosome 9% DMSO
Method: subtomogram averaging / : Chen JK, Cai S, Liu T, Ruan W, Ng CT, Shi J, Surana U, Gan L

EMDB-37198: 
The cryo-EM structure of type1 amyloid beta 42 fibril in AD2 patient.
Method: helical / : Zhao QY, Tao YQ, Liu C, Li D

EMDB-42948: 
Microtubule protofilament reconstruction in CCP5:microtubule class#1 complex
Method: single particle / : Chen J, Zehr EA, Gruschus JM, Szyk A, Liu Y, Tanner ME, Tjandra N, Roll-Mecak A

EMDB-44544: 
Microtubule protofilament reconstruction in CCP5:microtubule class#2 complex
Method: single particle / : Chen J, Zehr EA, Gruschus JM, Szyk A, Liu Y, Tanner ME, Tjandra N, Roll-Mecak A

EMDB-44545: 
Microtubule Protofilament reconstruction in CCP5:microtubule class#3 complex
Method: single particle / : Chen J, Zehr EA, Gruschus JM, Szyk A, Liu Y, Tanner ME, Tjandra N, Roll-Mecak A

EMDB-37170: 
The cryo-EM structure of type1 amyloid beta 42 fibril.
Method: helical / : Zhao QY, Tao YQ, Liu C, Li D
PDB-8kew: 
The cryo-EM structure of type1 amyloid beta 42 fibril.
Method: helical / : Zhao QY, Tao YQ, Liu C, Li D

EMDB-60197: 
Portal-tail of Vibrio cholerae typing phage mature VP1
Method: single particle / : Liu HR, Pang H

EMDB-60199: 
portal-tail of Vibrio cholerae typing phage release VP1
Method: single particle / : Liu HR, Pang H

EMDB-60702: 
Capsid of Vibrio cholerae phage mature VP1
Method: single particle / : Liu HR, Pang H

PDB-8zkk: 
Portal-tail of Vibrio cholerae typing phage mature VP1
Method: single particle / : Liu HR, Pang H

PDB-8zkm: 
portal-tail of Vibrio cholerae typing phage release VP1
Method: single particle / : Liu HR, Pang H

EMDB-43835: 
Structure of biofilm-forming functional amyloid PSMa1 from Staphylococcus aureus
Method: helical / : Hansen KH, Byeon CH, Liu Q, Drace T, Boesen T, Conway JF, Andreasen M, Akbey U
EMDB-38099: 
Cryo-EM structures of RNF168/UbcH5c-Ub in complex with H2AK13Ub nucleosomes determined by intein-based E2-Ub-NCP conjugation strategy
Method: single particle / : Ai HS, Tong ZB, Deng ZH, Pan M, Liu L
EMDB-38100: 
Cryo-EM structures of RNF168/UbcH5c-Ub/nucleosomes complex determined by activity-based chemical trapping strategy
Method: single particle / : Ai HS, Tong ZB, Deng ZH, Pan M, Liu L
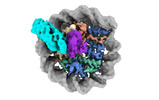
EMDB-38101: 
Cryo-EM structures of RNF168/UbcH5c-Ub in complex with H2AK13Ub nucleosomes determined by activity-based chemical trapping strategy (adjacent H2AK13/15 dual-monoubiquitination)
Method: single particle / : Ai HS, Tong ZB, Deng ZH, Pan M, Liu L
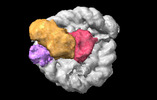
EMDB-38102: 
Cryo-EM map of RNF168/UbcH5c-Ub/nucleosome determined by E2-Ub-NCP conjugation strategy
Method: single particle / : Ai H, Zebin T, Deng Z, Pan M, Liu L

PDB-8x7i: 
Cryo-EM structures of RNF168/UbcH5c-Ub in complex with H2AK13Ub nucleosomes determined by intein-based E2-Ub-NCP conjugation strategy
Method: single particle / : Ai HS, Tong ZB, Deng ZH, Pan M, Liu L

PDB-8x7j: 
Cryo-EM structures of RNF168/UbcH5c-Ub/nucleosomes complex determined by activity-based chemical trapping strategy
Method: single particle / : Ai HS, Tong ZB, Deng ZH, Pan M, Liu L

PDB-8x7k: 
Cryo-EM structures of RNF168/UbcH5c-Ub in complex with H2AK13Ub nucleosomes determined by activity-based chemical trapping strategy (adjacent H2AK13/15 dual-monoubiquitination)
Method: single particle / : Ai HS, Tong ZB, Deng ZH, Pan M, Liu L

EMDB-60397: 
EcGK filament at APO state. (CASP target)
Method: single particle / : Zhang T, Leng Q, Liu LJ

EMDB-60066: 
Cryo-EM structures of RNF168/UbcH5c-Ub in complex with H2AK13Ub nucleosomes (two Ub conformation)
Method: single particle / : Ai HS, Tong ZB, Deng ZH, Tian CL, Liu L, Pan M
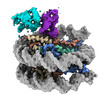
EMDB-39800: 
cryo-EM map of RNF168(1-193) in complex with Ubc5c-Ub conjugated nucleosome at a resolution of 3.23 angstrom
Method: single particle / : Ai HS, Tong ZB, Deng ZH, Tian CL, Liu L, Pan M
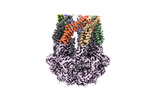
EMDB-36965: 
CryoEM structure of LonC protease hepatmer, apo state
Method: single particle / : Li M, Hsieh K, Liu H, Zhang S, Gao Y, Gong Q, Zhang K, Chang C, Li S
EMDB-36966: 
CryoEM structure of LonC protease open hexamer, apo state
Method: single particle / : Li M, Hsieh K, Liu H, Zhang S, Gao Y, Gong Q, Zhang K, Chang C, Li S
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search





 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
